Authors: Emily Hann, Debolina Majumdar, Daniel Layton, Mohamed Fareh, David M Cahill, Mark Ziemann, Beata Ujvari, Karel A Schat, Arjun Challagulla
Source: Functional & Integrative Genomics (Nov 2025)
Abstract
The CRISPR-Cas13 system has emerged as a powerful platform for programmable RNA targeting, offering efficient and sequence-specific silencing of coding and non-coding transcripts. The RNA-targeting capabilities of CRISPR-Cas13 have been harnessed to silence transcripts harbouring pathogenic mutations and combat infectious diseases. However, the molecular basis of on-target and collateral activity are not completely understood, limiting the utility of Cas13 systems.
In this study, we delineate the principles for the development of effective crRNAs by targeting DsRed fluorescence reporter and synthetic influenza mRNA in chicken fibroblast DF1 cells. To systematically determine the optimal design for RfxCas13d crRNA, we investigated the minimum length of the crRNA, importance of protospacer flanking sequence, degree of mismatch tolerance, and off target effects.
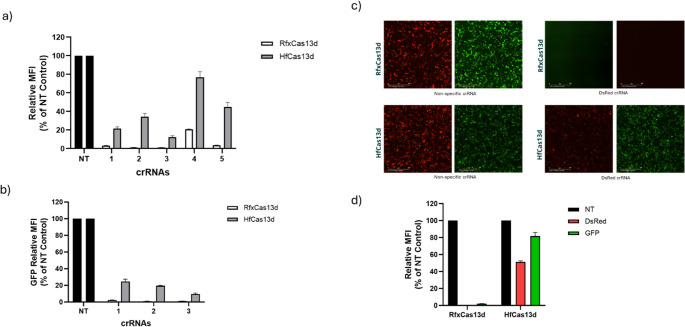
Our data reveal variable knockdown levels between crRNAs, in which several crRNAs achieved over 95% target knockdown. We show that crRNAs exhibit a high degree of tolerance to single-nucleotide mismatches, regardless of their position in the spacer sequence. However, 4-nt mismatches between the spacer and the target significantly reduces targeting efficacy, whereas eight nucleotide mismatches completely abolish the activity of RfxCas13d. Finally, we compared targeting efficiency and collateral activity of two widely used RfxCas13d and HfCas13d variants.
Our data extend current understanding of Cas13d-mediated RNA targeting and offer a framework for rational crRNA design to enhance effectiveness in diverse applications, including antiviral strategies.
Discover more from The Wild Genes Group
Subscribe to get the latest posts sent to your email.